What You're Looking At
This analysis covers eight Gulf species plus and two "True Trout" species. From the Gulf, four belong to Sciaenidae (the drums and croakers): Red Drum, Black Drum, Speckled Trout, and Atlantic Croaker. The remaining bony fish are Southern Flounder (Paralichthyidae), Sheepshead (Sparidae), and Flathead Mullet (Mugilidae). Bull Shark is the outgroup. Rainbow Trout and Brown Trout were added to directly test whether Speckled Trout has any evolutionary relationship to true trout.
Three COI1 sequences per species were downloaded from GenBank (24 total), then aligned with MUSCLE7 to a ~660 bp matrix. Multiple sequences per species act as a quality check: if all three cluster tightly together, the data are clean. Every within-species group clustered perfectly here. Trees were built with NJ4, ML5, and Bayesian inference6.
Before looking at the tree, where would you predict Speckled Trout clusters based on its common name? Was your prediction right?
Most people predict it clusters with Rainbow and Brown Trout. YOU WOULD BE SO WRONG! Its inside Sciaenidae alongside Red Drum and Atlantic Croaker at 100% support. Common names here reflect appearance and taste.
The Neighbor-Joining Tree *Note* I Need to edit the trees code to include bootsrtap vals
I am missing values on the interactive tree
The NJ tree uses pairwise K803 distances. Results show that Rainbow Trout and Brown Trout cluster together at 100% bootstrap, completely separate from every Gulf fish. Speckled Trout sits inside Sciaenidae with Red Drum, Black Drum, and Atlantic Croaker at 100% support. The branch separating Salmonidae from Sciaenidae is one of the longest in the tree.
Within Sciaenidae, Red Drum and Black Drum are sister taxa — natural hybrids between them are documented. Southern Flounder sits isolated from all Perciformes at 100% bootstrap; its flat body evolved independently, consistent with the polyphyletic origin of the flatfish body plan.13 Internal Sciaenidae nodes show 30–47% bootstrap; the family itself is at 100%, but COI alone can't resolve the exact branching order within it.
Internal Sciaenidae nodes show 30–47% bootstrap. Does that mean the family grouping is in doubt?
No. The node uniting all four Sciaenidae is at 100%. Low internal support only means we can't resolve the exact order in which those four species diverged. The family is well-supported.
In the outgroup explorer, switch between Bull Shark and Speckled Trout as outgroup. What changes, and what stays the same?
With Bull Shark, the root falls between all bony fish and the shark: correct. With Speckled Trout, the root is forced inside Sciaenidae, making the drum family look split, the underlying data does not change. Rainbow and Brown Trout still cluster together regardless of root choice. Outgroup choice changes direction, not topology.
Interactive: Choose Your Outgroup
Select any species as the evolutionary outgroup and see how the tree changes. Some choices are biologically justified; others are instructive mistakes.
Trees are Neighbor-Joining (K80 distances), generated in R using the ape package. Each tree uses the _1 sequence of the selected species as outgroup. Cladogram style; branch lengths not drawn to scale. Rainbow Trout and Brown Trout (Salmonidae) are included to anchor the true phylogenetic position of “trout” relative to Gulf fish species.
The Maximum Likelihood Tree
The ML tree used GTR+Gamma+I : all six substitution rates estimated independently, rate variation across sites modeled with a gamma distribution, and invariant sites accounted for. The family-level topology came out identical to NJ. Salmonidae holds at 100% bootstrap, cleanly separated from Sciaenidae, and Speckled Trout falls within Sciaenidae at the same support. Internal Sciaenidae nodes sit at 25–64%; the same uncertainty seen in NJ, which suggests this reflects genuine signal rather than a modeling artifact. Three sequences (Bull Shark 3, Southern Flounder 3, Flathead Mullet 2) shift position slightly, though their within-species distances are ≤0.0016, leading us to belive that placement is confident regardless of method.
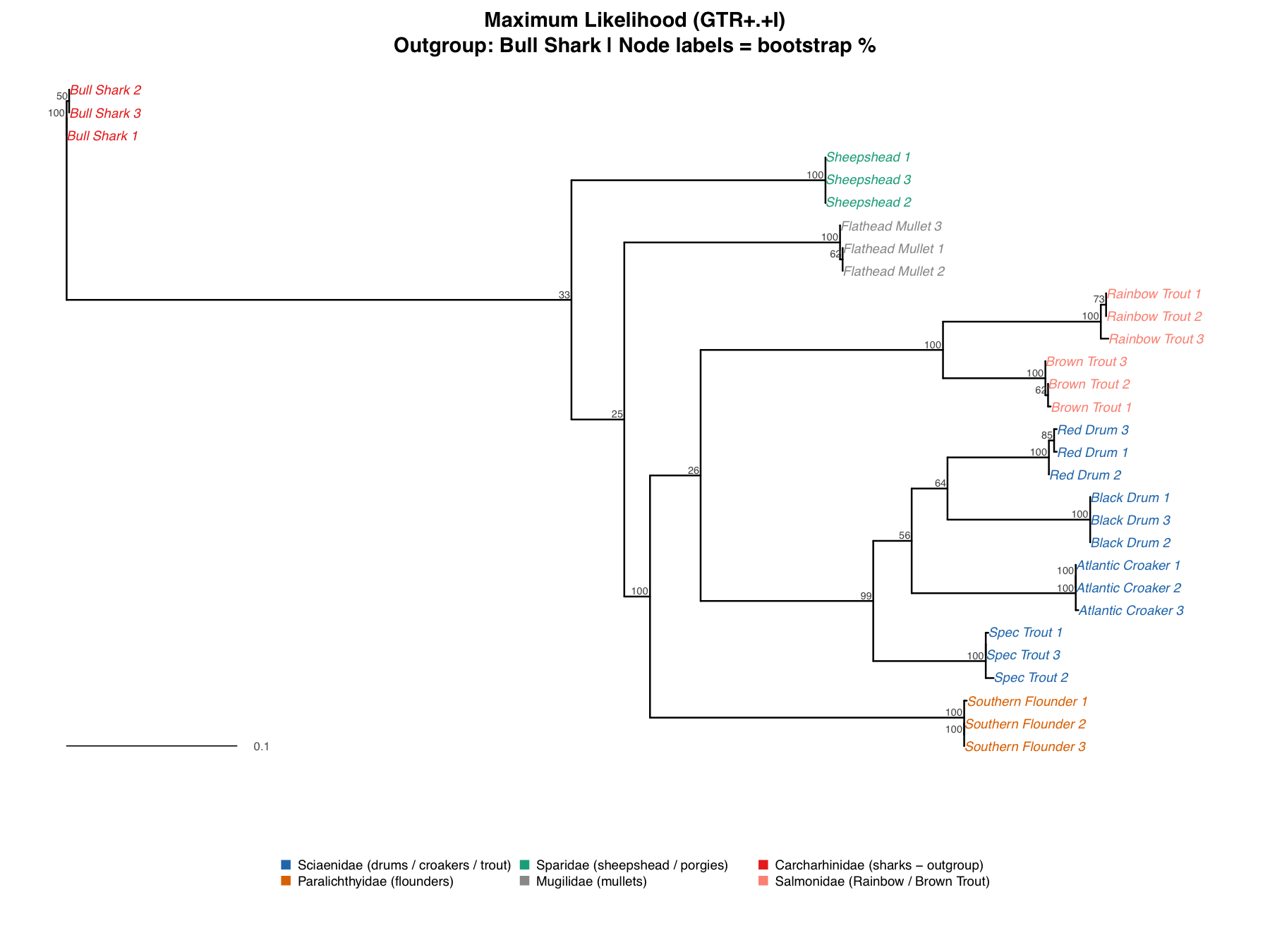
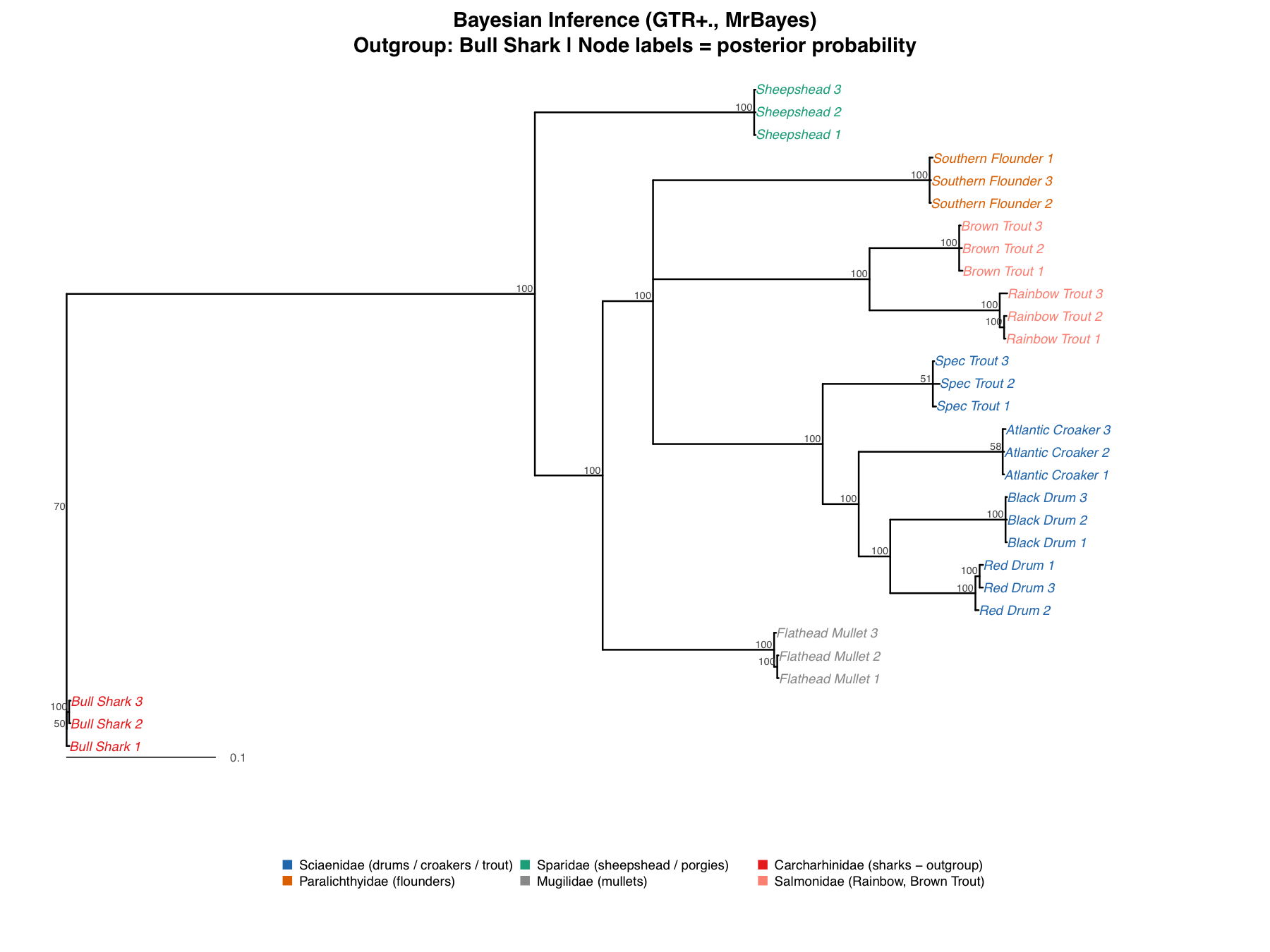
The Bayesian Tree
The Bayesian analysis (MrBayes, GTR+Gamma, 500,000 MCMC generations) gives the cleanest result. Nearly every node is PP = 1.00, including the Sciaenidae node, the Salmonidae node, and the node separating the two families. Speckled Trout is at PP = 1.00 inside Sciaenidae. The DNA is unambiguous: it's a drum, not a trout.
The only lower-support nodes (PP = 0.70, 0.58, 0.51) are internal splits within Sciaenidae, the same ones that showed weak support in NJ and ML. All three methods agreeing on where uncertainty lies confirms this is a biological limitation, not a methodological one. A posterior probability of 0.58 doesn't mean the node is wrong; it means there's 58% probability for that branching order and 42% for alternatives. PP values tend to run numerically higher than ML bootstrap and aren't directly comparable.
PP = 0.58 at one Sciaenidae node. A student says the tree is probably wrong there. Correct them.
PP = 0.58 means 58% probability for that branching order, 42% for alternatives among those same taxa. The family grouping is at PP = 1.00 and not in doubt. The uncertainty is only about which drum diverged first, a much narrower question that needs more data to resolve.